LVE Tutorial
Likhtman-McLeish theory
Start RepTate and create a new LVE Application
 :
:
Drag and drop a file(s) with a
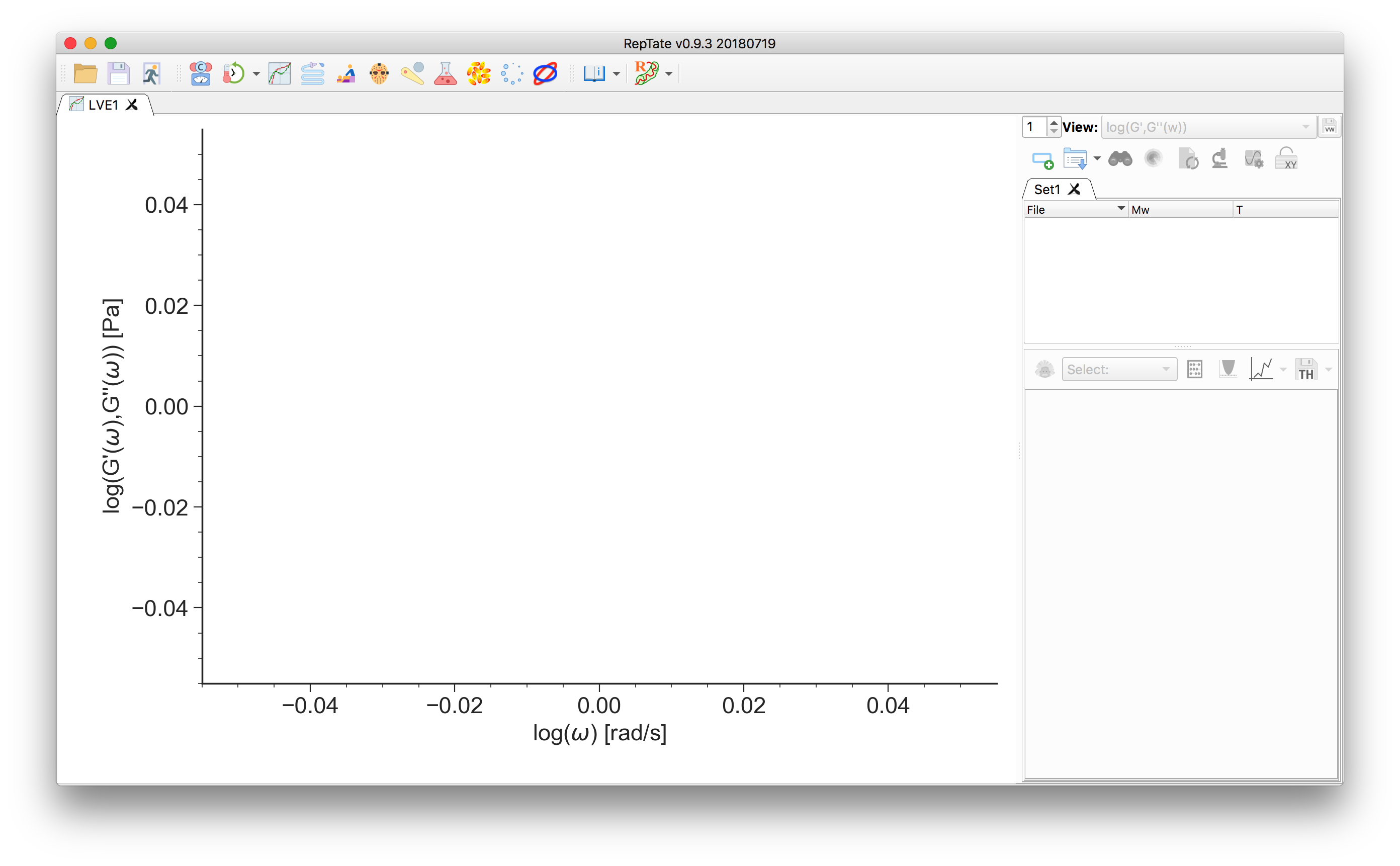
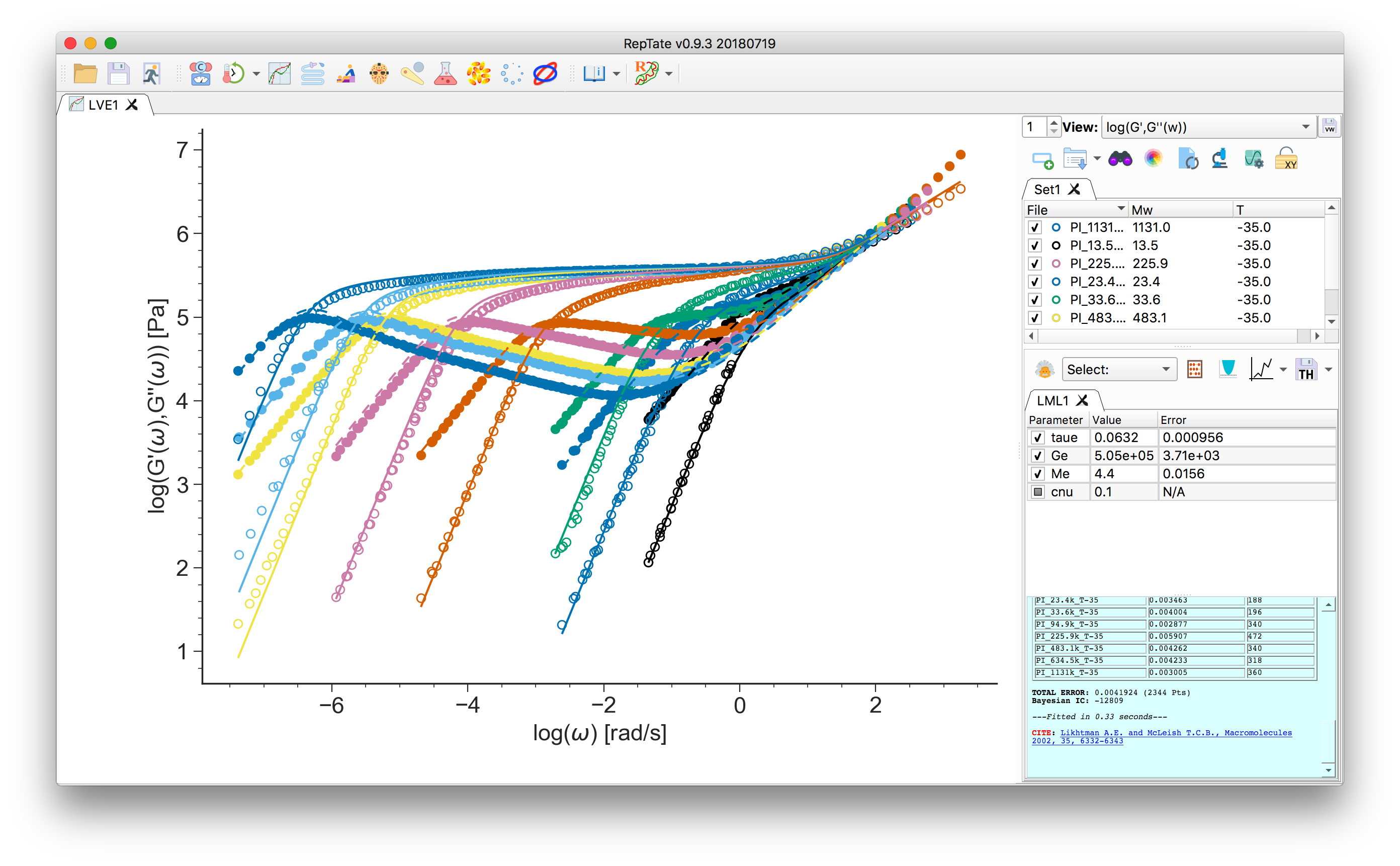
.ttsextension, e.g. the set ofPI_**k_T-35.ttsfiles in thedata/PI_LINEAR/folder, with molecular weight from “13.5k” to “1131k”. See Data Files for a description of the data file organization.
Select a theory, e.g. “Likthman-McLeish”
 , and press
, and press  to create it (calculation is done with default parameter values).
Press “Minimize Error”
to create it (calculation is done with default parameter values).
Press “Minimize Error”  .
.
BOB LVE predictions
BOB LVE uses a “polyconf” file, which contains information on the polymer architectures.
The “polyconf” files must carry a .dat. They can be generated by theories in the React application in RepTate
or by BOB software (Chinmay Das and Daniel Read) outside RepTate (https://sourceforge.net/projects/bob-rheology/files/bob-rheology/).
Start RepTate and create a new LVE Application
 :
:
Drag and drop a file with a
.ttsextension (see Data Files) OR add a dummy file if you do not have data corresponding to the “polyconf” file by clicking , which brings the following dialog.
, which brings the following dialog.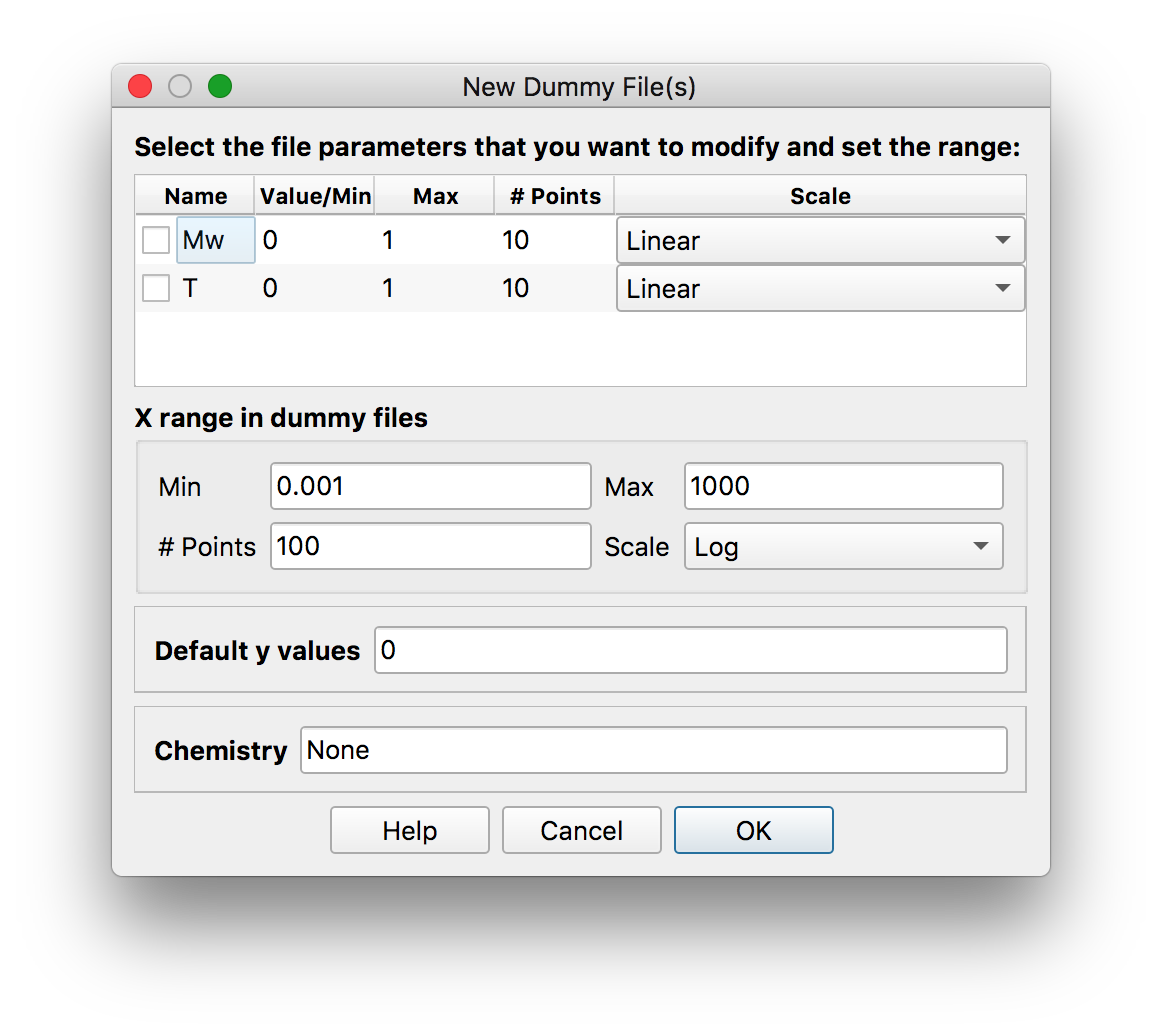
Note
The xrange of BOB predictions will correspond exactly to the xrange of the (dummy) file.
Select BOB theory,
 , and press
, and press  to create it. Then press “Calculate”
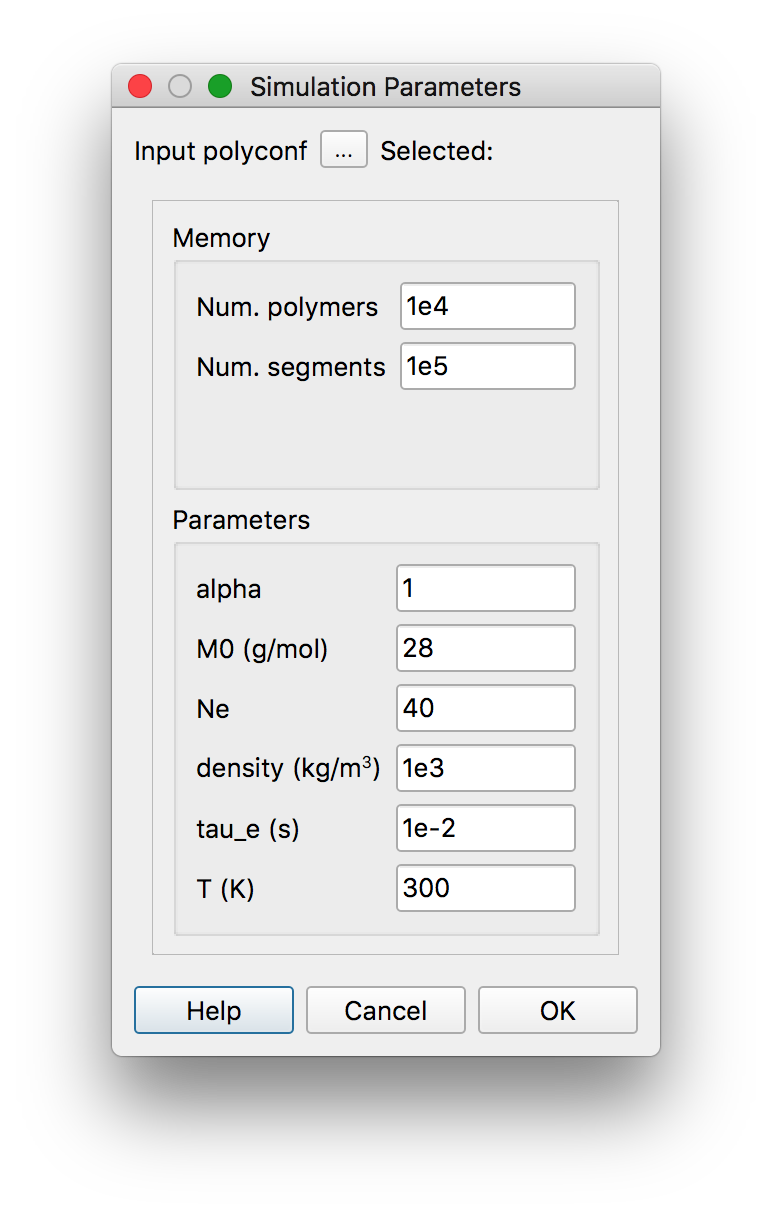
to create it. Then press “Calculate”  . The following dialog
appears, where, from top to bottom, we can select:
. The following dialog
appears, where, from top to bottom, we can select:the “polyconf” file (with a
.datextension)the maximum number of polymers
the maximum total number of arms (segments attached to a branch point)
alpha: the tube dilatation exponentM0: the molar mass of a monomerNe: the number of monomers in an entanglement segmentthe mass density of the polymer
the entanglement relaxation time
the temperature
Hint
Click the “Help” button to show the original documentation of BOB v2.3 by Chinmay Das.
Click “OK” to launch the calculations of the linear rheology. In the theory text-box, you will find informations related to BOB simulation parameters and to the topology of the polymers found in the “polyconf” file.