Creep: General description
Purpose
Data Files
The first line of the file should contain the sample parameters separated by semi-colons (
;). It may contain any number of parameters which will be read and saved as file-parameter in RepTate. In unit-aware applications, a file parameter value may also include an explicit unit, for exampleMw=1131 Da;T=25 ºC;gdot=0.1 1/s;.Then the data columns should appear, separated by spaces or tabs.
In unit-aware applications, column headers may include units in square brackets or parentheses, for example
t [min]orG' [kPa]. See Units for the supported units and current limitations.
.creep extension
Text files with .creep extension should be organised as follows:
.creepfiles should contain at least the parameter value for the:Applied stress
stress
2 columns separated by spaces or tabs containing respectively:
time, \(t\),
strain, \(\gamma\),
A correct .creep file looks like:
stress=10;chem=PE;label=CM3;T=150;
t strain stress T
s - Pa C
1E-3 1.413E-5 10 149.9
1E-3 -9.419E-6 10 149.9
1E-3 -2.826E-5 10 149.9
... ... ... ...
Views
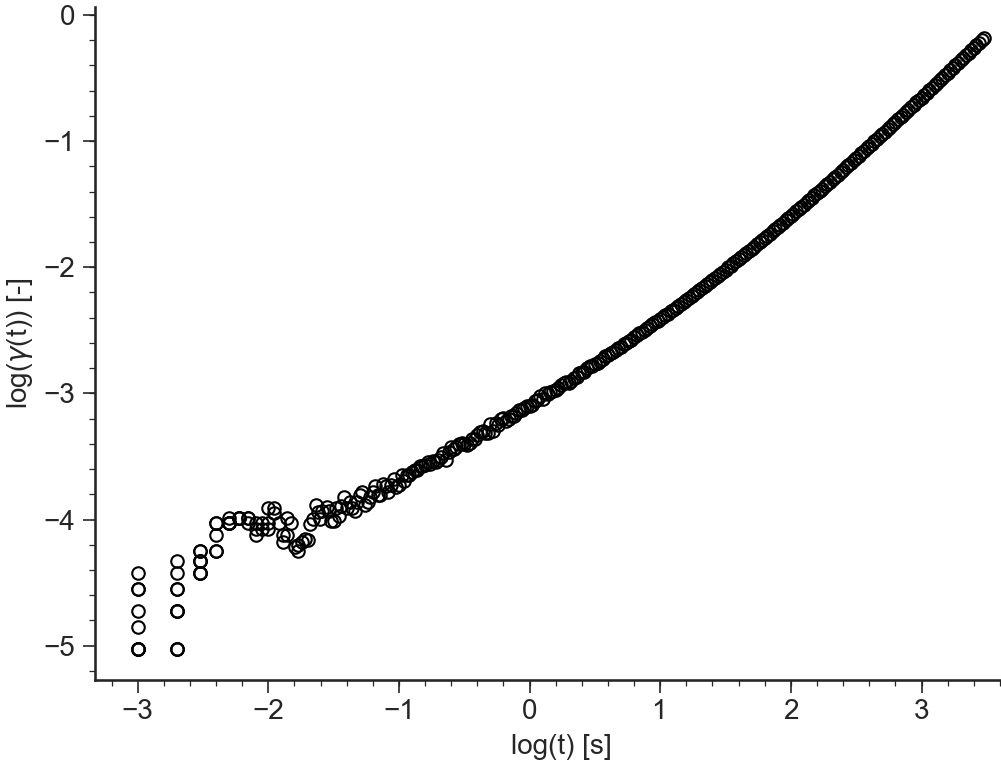
log(gamma(t))
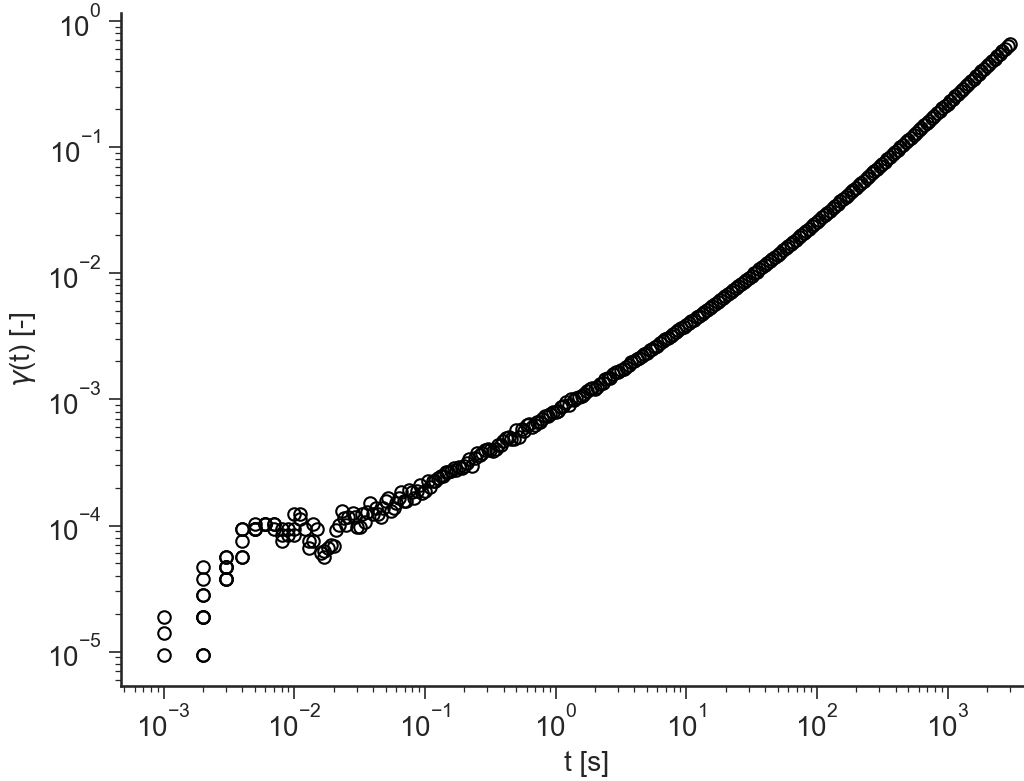
gamma(t)
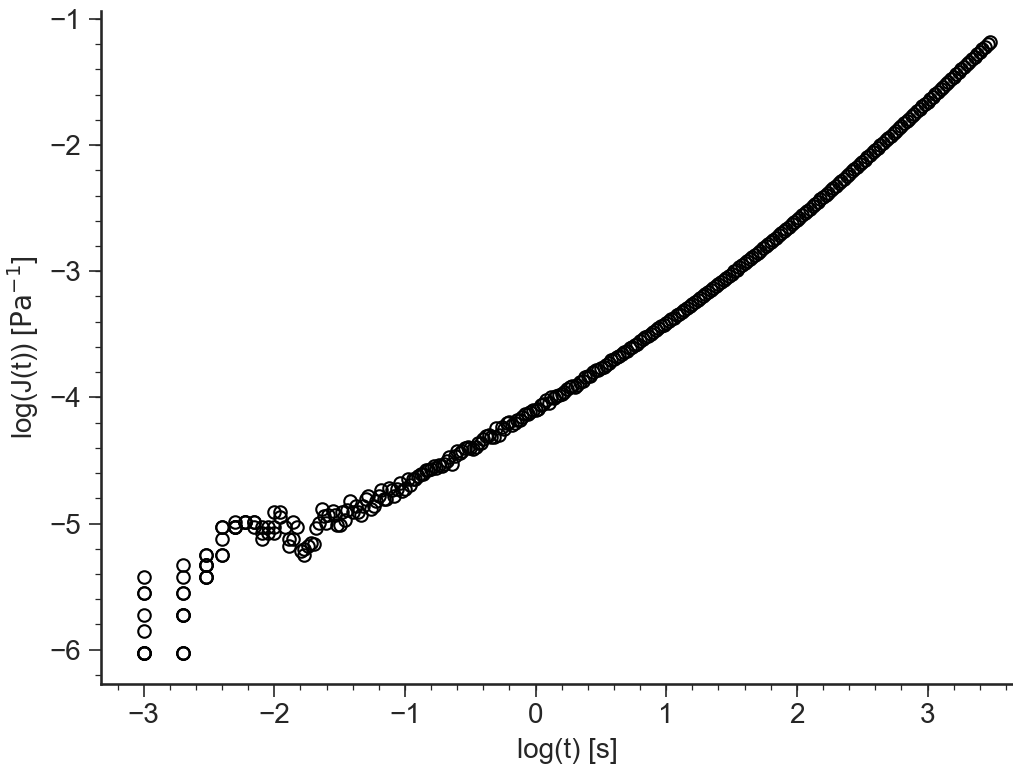
log(J(t))
J(t)
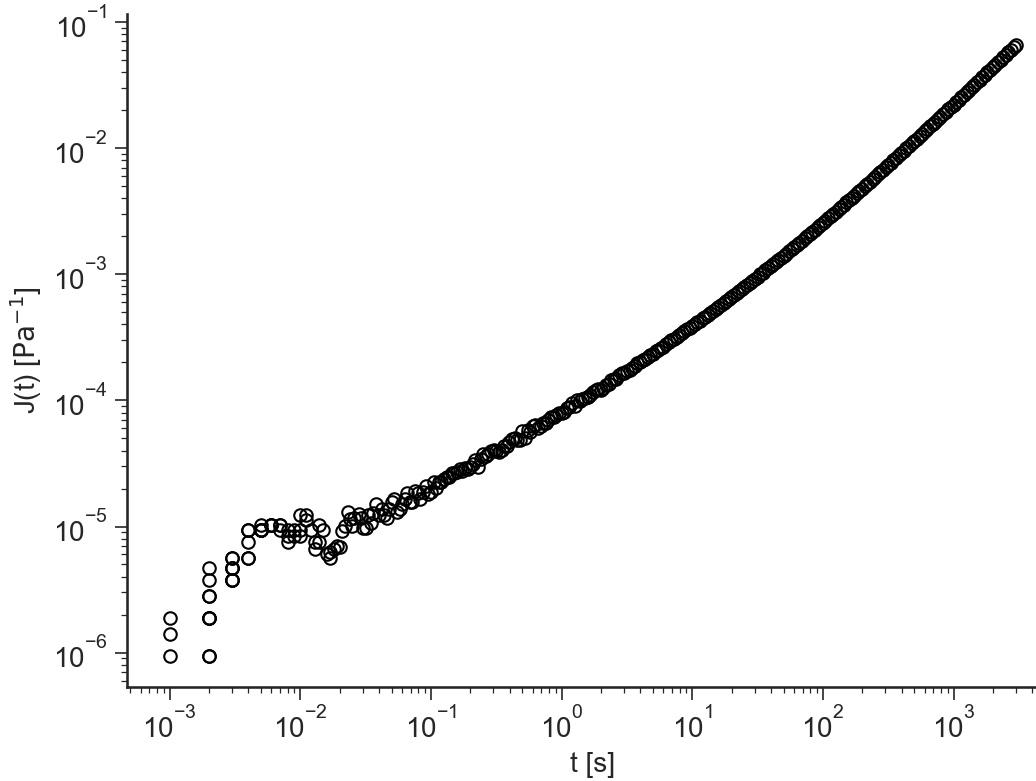
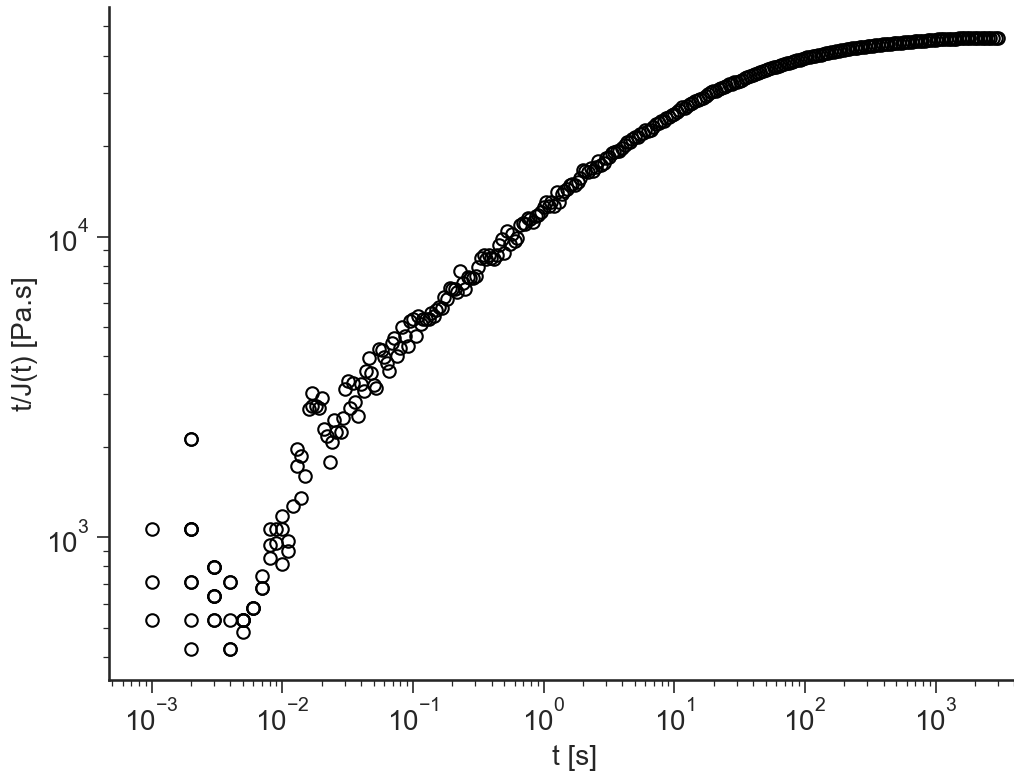
t/J(t)